

and thair research arias

Computational Biology

Metagenomics

Metabolic Engineering

Computational Biology
Rising costs and a depleting supply of oil, as well as environmental concerns, have led to strong interest in renewable fuels and chemicals.In the last decade, there was considerable interest for microbial production of biofuels by metabolic engineering approach as an attractive alternative to transportation fuels. Recent advancements of synthetic biology have led to the further development of genetic engineering of microorganism, which has been great motivation for developing strategy for microbial production of biofuels. Rational design of metabolism is very important for production of biofuels. In silico prediction of metabolic flux distribution of the metabolic pathways have enabled us to decide the time consuming steps in metabolic engineering. In recent years significant efforts have been made to engineer microorganisms to produce bioethanol, higher chain alcohols, fatty acids and isoprenoid based biodiesel. Metabolic engineering involves improvement of biofuels formation through the modification of specific genes or addition of new genes involved in biochemical reactions with the use of recombinant DNA technology. System-level approach to analyze and engineer metabolism based on flux distributions obtained from metabolomics and 13C metabolic flux analysis have been extensively used by our group to produce biofuels in Escherichia coli and yeast.
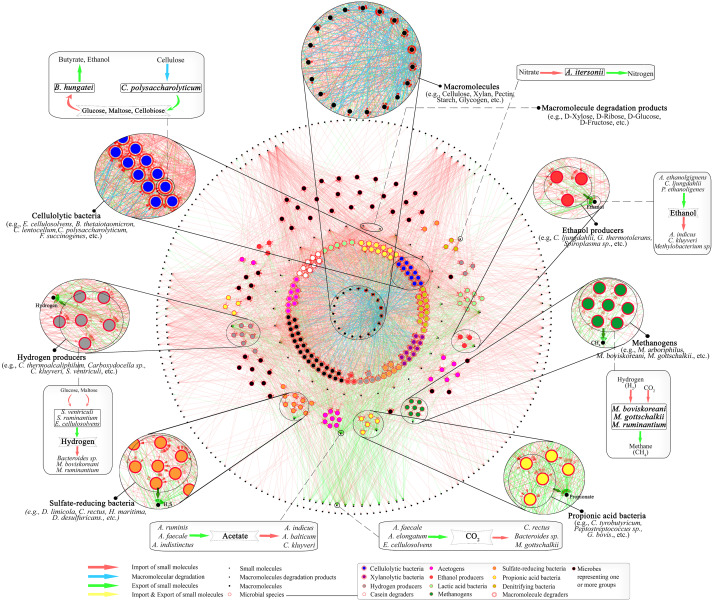
Diverse environmental condition reinforce the growth of different microbial communities with specialized characteristic. Metagenomic study along with computational technique disclose the structural and functional complexity of microbial niche. Culture independent approach of metagenomics helps to conquer the hurdle of characterization of unculturable microbes, enhancing clarity of community based microbiology. Gut microbiota has been one of the most fascinating area of research during several decades because of enormous microbial diversity and their metabolic potential. We are interested to study the cumulative potential of different gut microbiota that effectively translate the lignocellusoic biomass to biofuel. A system level understanding of the microbial interdependency and cross communication will decrypt the natural system lignocellulose degradative system leading towards the designing of a successful invitro mechanism for effective biomass conversion for biofuel production. Systematic mathematical model development of inter species interaction will help to identify key-species (influencer) in microbial communities having powerful lignocellulolytic property.
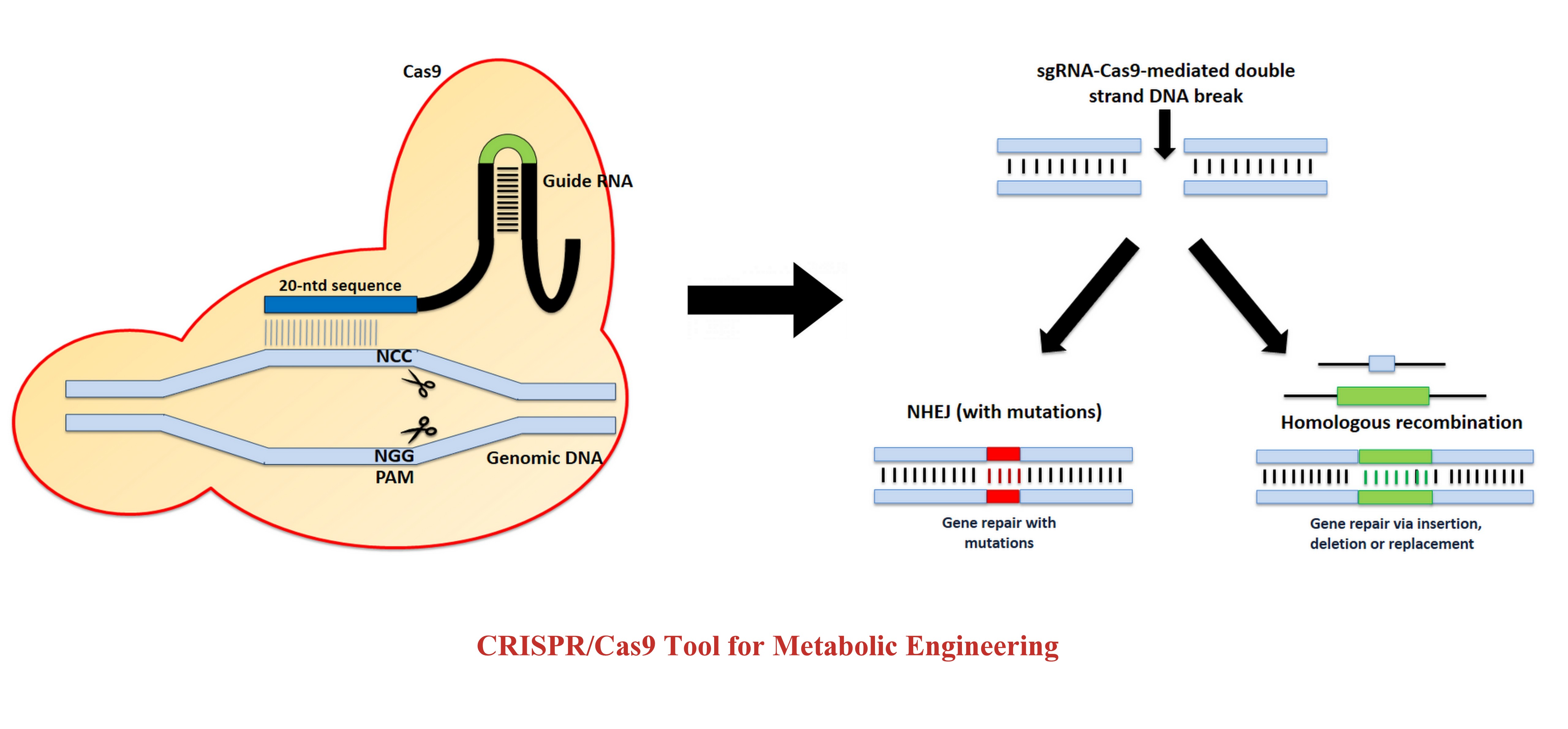
Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and CRISPR-associated (Cas) proteins were discovered in prokaryotes as an adaptive immune system against viral and plasmidal genome invasion. The type II CRISPR/Cas system uses a tracrRNA:crRNA hybrid to bind to Cas9 protein and guide it to create a site-specific double stranded break on target templates.Modification of the tracrRNA:crRNA hybrid to a single guide RNA that has a customizable recognition sequence to guide DNA cleavage has become a very popular technique in gene editing. CRISPR/Cas9 technique also allows a marker-less genome manipulation that enables several modifications on the same strain. Post-cleavage of DNA with CRISPR/Cas9, the DNA repair occurs via non-homologous end joining (NHEJ) or homologous recombination (HR). An increasing global energy demand and environmental concerns associated with petroleum fuels have necessitated the need for alternative liquid fuels (biofuels). Microbes such as yeasts can utilize sugars (generally hexoses) to produce different compounds such as free fatty acids (FFA) and its derivatives such as fatty-acid ethyl ester (FAEE), fatty alcohols and alkane/alkene. These biomolecules have energy content comparable to diesel and petrol, making them an important candidate for biofuels.

Molecular Dynamics Simulation is a classical mechanics based computational approach that enables to investigate the behaviour of a microsystem consisting a large number of interacting particles and to predict the bulk properties of the macrosystem at molecular level. GPU accelerated high-performance computing has revolutionized the theoretical approached to study the structure and dynamics of biological macromolecules from a nanosecond (ns) to milliseconds (ms) timescale in recent years. Our lab is particularly interested to explore the molecular mechanism of various biological processes in the domain of carbohydrates as well as protein chemistry using MD simulation. Currently, we are focussing to understand the dissolution of cellulose in Ionic liquid (1-Ethyl-3-methylimidazolium acetate) and water mixtures for designing a cost-effective and optimized lignocellulosic biomass pretreatment process using IL. We are also working to develop theoretical insights behind the IL tolerance of thermostable cellulolytic enzymes, which will guide the experimental biologists for engineering the existing cellulase with improved saccharification kinetics. Moreover, molecular simulations of proteins reveal multiple conformational states in a trajectory ensemble which can explain its in-vivo functionality. Steered Molecular Dynamics are also routinely carried out in our lab to study the force induced unfolding of proteins.
School of Energy Science and Engineering Sir J.C. Bose Laboratory Complex, West Bengal
+91-3222-260804
amitghosh@iitkgp.ac.in